Dan Kortschak
Software Engineer
Biography
Dan is actually doing science.
- Software engineering
- Graphical analysis
- Data visualisation
PhD in Genetics, 1998
The University of Adelaide
GDip Education, 2006
The University of Adelaide
BSc (Hons) in Biochemistry, 1993
The University of Adelaide
Projects
Projects I am involved in or own.

dex
dex is an El Gato Stream Deck controller system.

Ugg Boot
Ugg boot provides a simple way to update Go executables and list available versions using module version information embedded in the executable.

rgo
rgo is an R/Go integration tool.

mbg
mbg is an mbox forensics tool written for fun.

AusOcean
AusOcean is an environmental not-for-profit with a mission to help our oceans through the use of technology.
typo
The typogenetics game from Gödel, Escher, Bach: an Eternal Golden Braid.
arrgh
arrgh is a Go interface to the OpenCPU R server system.

bíogo
bíogo is a collection of packages for bioinformatics written in the Go programming language.

ev3go
ev3go is a Go interface to LEGO® Mindstorms-compatible robotics platform built on the ev3dev.org Linux distribution.

Gonum
Gonum is a collection of scientific software packages written for the Go programming language.
utter
utter implements a deep pretty printer for Go data structures.
zalgo
Invoke the hive mind.
Selected Publications
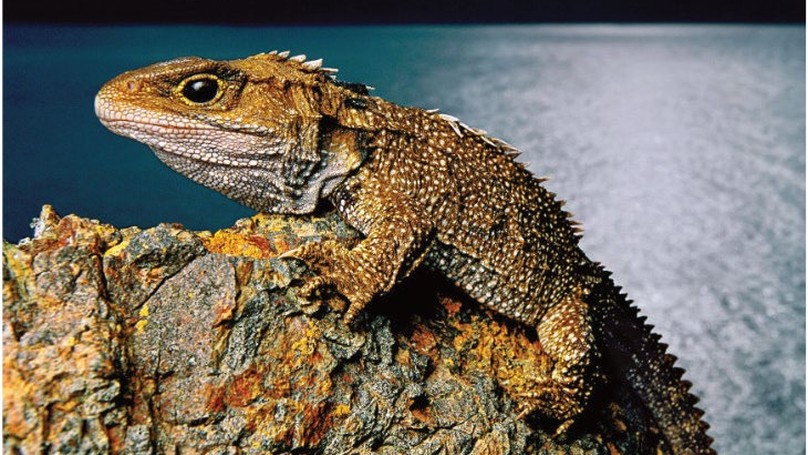

The tuatara (Sphenodon punctatus)—the only living member of the reptilian order Rhynchocephalia (Sphenodontia), once widespread across Gondwana—is an iconic species that is endemic to New Zealand. A key link to the now-extinct stem reptiles (from which dinosaurs, modern reptiles, birds and mammals evolved), the tuatara provides key insights into the ancestral amniotes. Here we analyse the genome of the tuatara, which—at approximately 5 Gb—is among the largest of the vertebrate genomes yet assembled. Our analyses of this genome, along with comparisons with other vertebrate genomes, reinforce the uniqueness of the tuatara. Phylogenetic analyses indicate that the tuatara lineage diverged from that of snakes and lizards around 250 million years ago. This lineage also shows moderate rates of molecular evolution, with instances of punctuated evolution. Our genome sequence analysis identifies expansions of proteins, non-protein-coding RNA families and repeat elements, the latter of which show an amalgam of reptilian and mammalian features. The sequencing of the tuatara genome provides a valuable resource for deep comparative analyses of tetrapods, as well as for tuatara biology and conservation. Our study also provides important insights into both the technical challenges and the cultural obligations that are associated with genome sequencing.
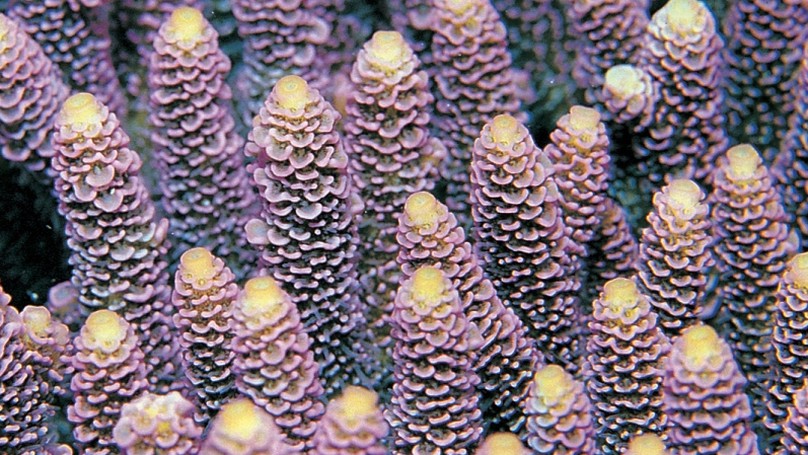

A significant proportion of mammalian genes are not represented in the genomes of Drosophila, Caenorhabditis or Saccharomyces, and many of these are assumed to have been vertebrate innovations. To test this assumption, we conducted a preliminary EST project on the anthozoan cnidarian, Acropora millepora, a basal metazoan. More than 10% of the Acropora ESTs with strong metazoan matches to the databases had clear human homologs but were not represented in the Drosophila or Caenorhabditis genomes; this category includes a surprising diversity of transcription factors and metabolic proteins that were previously assumed to be restricted to vertebrates. Consistent with higher rates of divergence in the model invertebrates, three-way comparisons show that most Acropora ESTs match human sequences much more strongly than they do any Drosophila or Caenorhabditis sequence. Gene loss has thus been much more extensive in the model invertebrate lineages than previously assumed and, as a consequence, some genes formerly thought to be vertebrate inventions must have been present in the common metazoan ancestor. The complexity of the Acropora genome is paradoxical, given that this organism contains apparently few tissue types and the simplest extant nervous system consisting of a morphologically homogeneous nerve net.
Teaching
I have written and taught a variety of courses in bioinformatics and software engineering practice for scientists, and coordinated the Master of Bioinformatics programme at the University of Adelaide from 2018 to 2020.
A half day course in basic git use is available here.
Other interests
Dan has other interests outside science and software development. He will share these things face to face.
